Haematopoiesis relies on the ability of hematopoietic stem cells to progress through a systematic hierarchy to produce lineage-restricted progenitors that terminally differentiate into phenotypically distinct types of mature hematopoietic cells. This process is precisely coordinated by the combinatorial activity of lineage-specifying transcription factors (TFs). Indeed, the critical transcriptional program of every hematopoietic cell type, and indeed of all cell types throughout the body, requires a set of core TFs for its proper execution.
A frequently-overlooked component of the cellular transcriptional program is the transcription of ribosomal RNA (rRNA), the major component of ribosomes. Ribosomal RNA comprises 90% of total cellular RNA, and its transcription from hundreds of rDNA genes by RNA polymerase I is one of the most intense transcriptional processes in the cell. Different progenitor and mature cell types in the hematopoietic tree have different sizes, ribosome abundances, and rates of protein synthesis. Tight control of ribosome abundance is essential for a normal cellular proteome, and different cell types within the hematopoietic tree have varied rRNA transcription rates. However, there has been limited study of the molecular basis of lineage-specific regulation of rDNA transcription in hematopoiesis.
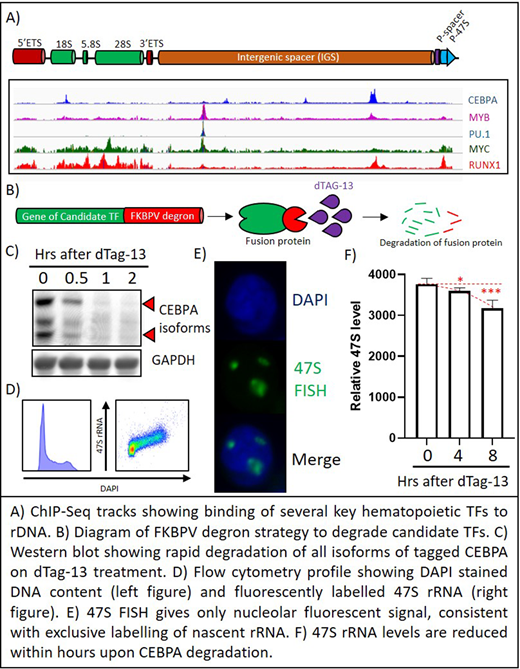
We explored the binding to rDNA of over twenty key hematopoietic TFs that were determined by the Broad Institute DepMap database to be crucial for survival of hematopoietic cell lines. Using a customized bioinformatics pipeline, we mapped over three hundred ChIP-Seq datasets for these factors (generated by us as well as publicly available from ENCODE and GEO) to mouse and human rDNA assemblies, and found that several essential hematopoietic TFs such as MYC, MYB, RUNX1, PU.1, CEBPA and others bind to rDNA at conserved sites and motifs (Fig A). MYC is well-known as a master regulator of rDNA, and RUNX1 was recently reported to bind rDNA, but most of the others have never been linked to rDNA, and their functional roles in regulating rDNA transcription have not been explored.
We picked for further study CEBPA, a crucial TF required for specification of granulocyte-monocyte progenitors (GMPs). For our experiments, we used the mouse HOXA9-ER cell line, which mimics GMPs. We used CRISPR/Cas9 and homologous recombination to fuse FKBPV degron into bi-allelic endogenous loci of the Cebpa gene in HOXA9-ER cells, and, upon addition of dTAG-13 (the ligand for FKBPV), the CEBPA-FKBPV fusion protein could be rapidly degraded within 2 hours (Fig B, C), providing us an experimental system to study the immediate consequences of CEBPA loss. In order to quantify the rate of rRNA transcription, we devised an assay titled "47S FISH-Flow" that combined fluorescent in-situ hybridization (FISH) using probes against nascent 47S rRNA with flow cytometry (Fig D, E). This assay not only allows us to quantify the rate of rRNA transcription on a per-cell basis in millions of cells, but also allows us to separately gate and quantify rRNA transcription in different stages of the cell cycle, eliminating a major confounder in bulk cell studies - cell cycle distribution. Using 47S FISH-Flow, we observed that degradation of CEBPA in the HOXA9-ER mouse GMP cell line led to decrease in synthesis of 47S rRNA within hours (Fig F) before any change in cell cycle or growth kinetics, and was followed by growth arrest in 24 hours.
In summary, we show that several critical hematopoietic TFs show abundant, conserved binding to rDNA, and the depletion of CEBPA rapidly reduces nascent rRNA, indicating that it directly promotes rRNA transcription. Our results, and the tools and experimental systems we have developed, shed light on an important and largely unexplored aspect of hematopoietic biology: the regulation of rRNA transcription by a wide range of lineage-specific hematopoietic TFs.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.